Introduction
Behind many botanical specimens lies a story, often glamourized, of plant species being torn from their wild origins. Rare plant prizes were eagerly sought after and taken from their native habitat to be propagated and sold in a booming horticultural trade that exploded following the age of exploration. Some plants suffered during this exploitation with resultant population decline. The plant trade was one of the earliest globalisation initiatives through which plants from all across the world were brought to and grown in botanic gardens. Now, some of the rarest of plants are held in public and private gardens across the world, some of these plants are even extinct in the wild. A review of rare plants in European botanic gardens collections showed approximately 54% of threatened plant species were in cultivation (Maunders et al., 2001). It is encouraging that so many are in cultivation, but what is the fate of these specimens? Do we admire them in pots and walled gardens or could we more actively utilise them for conservation? They harbour a genetic resource that could be used for conserving or even re-introducing populations.
A core issue in many conservation initiatives is to maximise diversity by increasing the number of individuals and thus increase the genepool. This is certainly the case in rare or threatened populations, although for certain unique populations, distinctiveness may be an important characteristic to maintain. In botanic gardens there tends to be only a few individuals in each collection, so the genepool is limited. One approach would be to pool individuals from many gardens into a breeding population and then propagate from this stock. However, even so, the origins of the material are often not known. In addition, the frequent practice of sharing and exchange of plants between gardens and a limited number of plant collectors will lead to a limited set of sources. This increases the risk of inbreeding and can negatively affect the fitness of a population (Charlesworth and Charlesworth, 1987). Molecular techniques offer potential tools for investigating the genetic diversity within a set of individuals or a population. They can be used to test for bias or restricted genepools in breeding programme populations. A small-scale pilot project was undertaken to test the use of DNA analysis in helping to determine the genetic basis of a plant collection of Delavay’s magnolia (Magnolia delavayi Franch.) in Irish gardens. The aim of the project was to determine the genetic base of a limited collection of M. delavayi and to test if this corresponds to historical records.
Objects and methods of research
The target plant – Magnolia delavayi
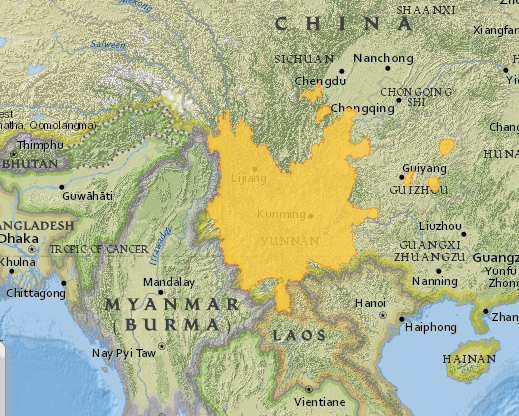
Fig. 1. A map of the natural distribution range of Magnolia delavayi (Map from IUCN).
Delavay’s Magnolia or the Chinese Evergreen Magnolia is a species of magnolia native to southern China. It was discovered to western science by the French missionary and plant collector, Père Jean-Marie Delavay. Delavay (1834-1895) was a plant collector for the French National Natural History Museum in Paris and contributed a large number of herbarium specimens to its collection (Kilpatrick, 2014). He also collected seed and fruit that were cultivated in the Jardin des Plantes in Paris. His collecting endeavours have been acknowledged in many plant names, M. delavayi being the most obvious. Magnolia delavayi was brought into cultivation by Ernest Henry Wilson in 1899 who made collections in China while employed by Veitch nurseries (Treseder, 1978). Although there are insufficient data to make recommendations regarding the current conservation status of the species (Cicuzza et al., 2007), Delavay’s magnolia was considered endangered until the most recent IUCN Red List was prepared in 2014. It is currently considered of “Least Concern” in the IUCN Red List (IUCN, 2015) due mainly to its large distribution range, which spreads across much of the Yunnan province and beyond (River & Wheeler, 2014). The distribution is mainly contiguous, although a few fragmented populations lie to the east and north of the Yunnan province (Fig. 1). Though this species is not under threat in the wild, it is a useful case to test the feasibility of the approach presented. The potential use of garden plants to re-introduce individuals or populations into the wild can be important for other threatened species. In order to do this it is important to understand the genetic base of the garden material.
Historical Context
The aim of the work was to assess if a set of specimens of Magnolia delavayi in Irish botanic gardens represent a single genotype or if they were multiple distinct genotypes. A preliminary historical search was undertaken to determine the origins of the material used, although data was not available for some specimens. Some information was available in Forrest (1984), but additional information was obtained from checking the records in the National Botanic Gardens, Glasnevin, Co. Dublin. There is a single living specimen in the National Botanic Gardens in Glasnevin and another in the arboretum in Kilmacurragh, Co. Wicklow. The specimen from Glasnevin is a donation from Veitch of Chelsea, England. There are records of two donations from Veitch in the register in Glasnevin, one in 1902 and another in 1908, but only one of these specimens survives today. Kilpatrick (2014) notes a sapling was sent from the Veitch nursery to the Royal Botanic Gardens, Kew in 1902 and that it was first exhibited by Veitch in October of 1912. There is a note in the Glasnevin records that the specimen in Glasnevin is from Yunnan, so it is likely that the specimen is from the main distribution area of M. delavayi rather than from the outlying populations (Fig. 1). The specimen in Kilmacurragh is a propagation from Glasnevin. Samples were also obtained from Mount Usher Gardens, in Co. Wicklow, Birr Castle in Co. Offaly and Lismore in Co. Waterford (Table 1). Six M. delavayi individuals were used and two additional samples of other Magnolia species (M. kobus DC. and M. grandiflora L.) were included as out-groups in the analysis (Table 1). There are two herbarium specimens of M. delavayi in the DBN herbarium at Glasnevin – one from the Royal Botanic Gardens, Kew, deposited by Elwes and Henry in 1911 as part of their research on The Trees of Great Britain and Ireland, and one from Birr, Co. Offaly deposited by Brian Morley in 1975 (Fig. 2).
Table 1. Samples used in the analysis
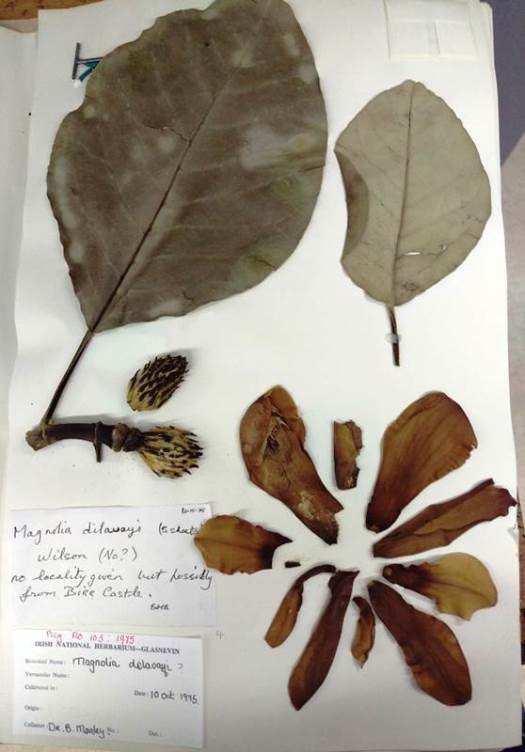

Fig. 2. A specimen of Magnolia delavayi in the DBN herbarium, Glasnevin.
Laboratory Methods
Leaf samples were stored in silica gel until DNA extraction was performed. DNA was extracted from the samples using QIAGEN DNeasy kits according to the manufacturer’s instructions. DNA was stored at -20° C until used and has been deposited in the DNAbank in Glasnevin.
Investigations were undertaken using RAPD (Random Amplified Polymorphic DNA) genetic fingerprinting and DNA sequencing. RAPD is a fingerprinting technique used to scan for variation across the entire genome, however, it has been criticised over its reproducibility (Jones et al, 1997). DNA sequencing is a more reliable and transferable technology. Specific DNA regions of the chloroplast genome were sequenced to assess variation. As this was a pilot study, only four regions were selected for testing. There were no published reports on genetic variation in M. delavayi, with the exception of the description of two flowering forms – the most common being a white form and the other being a red form (Xun et al., 1998). No reports of genetic diversity at the molecular level exist, so universal markers were utilised. Four regions were screened for DNA sequence variation; atpB-rbcL (Terachi, 1993), psbA-trnH (Kress et al., 2005), psbD-trnT (Shaw et al., 2007), trnL-F (Taberlet et al., 1991). PCR was performed using Bioline MyTaq mix on an Eppendorf Mastercycler. For details of the experiments contact the author. PCR products were cleaned with SureClean, Bioline, using the manufacturer’s protocol and were sent for sequencing using Macrogen, Korea. Sequences were aligned using Sequencher 4.10. Differences in the samples were calculated based on percentage differences across the entire sequences.
Results and discussion
The genetic fingerprinting approach using RAPD (Random Amplified Polymorphic DNA) analysis showed differences in the genetic fingerprint of sample M5 (Birr) compared with the other Irish samples and also showed the species outgroups to be clearly different from each other. However, it was decided to focus on the DNA sequencing data instead as RAPD results have been shown to be difficult to reproduce across labs (Jones et al., 1997).
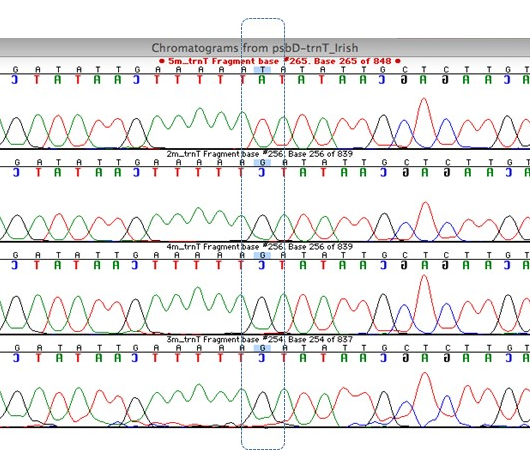
Fig 3. A screen-shot from a segment of the DNA alignment of samples M2-M5 showing DNA sequence variation in sample M5.
The four regions sequenced gave a total of 3068 bp of DNA sequence in eight specimens. The sequencing results showed clear differences between the species (M. delavayi to M. kobus and M. grandiflora), which is to be expected, but it also showed there was some variation between sample M5 (Birr) and all the other M. delavayi samples (Fig. 3, Table 2). The Birr sample differed in two locations in one of the regions analysed. The region in Fig. 3 shows variation at a single nucleotide, a SNP (Single Nucleotide Polymorphism). Regardless of a physical manifestation of this DNA variant, it allows us to distinguish genotypes as distinct and possibly different lineages.
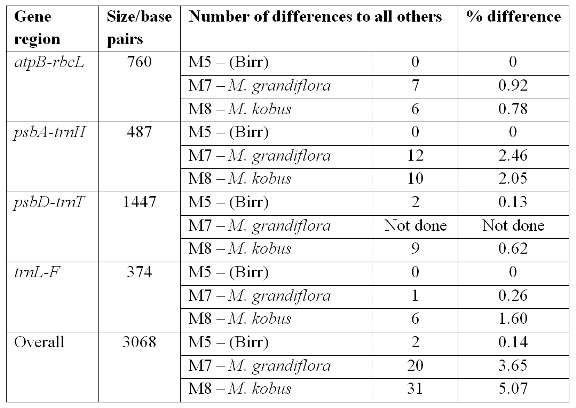
Although the difference is small between the Magnolia delavayi samples (0.14% over all the data), it does represent a clear difference between the genotypes in the Irish gardens. The results from the genetic fingerprinting technique were not used in the final analysis but they also distinguished the Birr M. delavayi sample from the others. The fingerprinting results also point to the potential for more variation rather than just one gene region. The species difference (M. kobus or M. grandiflora to M. delavayi) was also small – averaging between 3% and 5% across all the regions screened, so the individual difference is approximately 30 times smaller than a species difference. Magnolia grandiflora had a total of 20 differences and M. kobus a total of 31 differences. The Birr sample had two differences to the other M. delavayi samples. The DNA sequence results indicate that the Birr sample is unique and possibly from a different source location than that of the Glasnevin and other samples. In an untitled catalogue of trees from Birr dated around 1936 in the National Botanic Gardens archives, one of the Birr M. delavayi samples is noted as coming from Hilliers, England. However, a published list indicates the Birr specimen is from Veitch, as is the Glasnevin sample (Forrest, 1985). So, although the historical records are uncertain, the DNA evidence indicates the Birr Castle specimen is from a different source than that of Glasnevin, Kilmacurragh, Mount Usher and Lismore.
Table 2. Variation in the DNA regions sequenced
Conclusions
This pilot study revealed two important findings. First, it identified a variable region that can be used to distinguish different individuals of Magnolia delavayi and second it showed that the specimens in Irish botanic gardens contain at least two different genotypes. Out of four regions screened the psbD-trnT region was the only one that showed variation, but it showed two variable sites across the individuals. This region can be tested on other magnolia species. One particular species that would be a worthwhile subject is Magnolia stellata (Siebold & Zucc.) Maxim. This species is commonly available in cultivation but is endangered in the wild (IUCN, 2015).
The analysis undertaken in this project was limited and there is a need to progress it further to assess the potential utility of garden specimens in future conservation programmes. Further work could entail using other molecular markers on a greater array of samples from living collections. The study should be broadened to a worldwide assessment. It would be especially useful to compare the garden specimens against wild populations, in particular against marginal populations outside of the main Yunnan province distribution of M. delavayi.
While gardens may have limited numbers of individuals in a collection, a relatively quick and easy screen of genetic diversity can be used to focus efforts in the right direction for conservation purposes. Botanic gardens have a long and fruitful history of contributing to scientific investigation and in particular to understanding biology. The use of DNA analysis in aiding conservation is another welcome addition and one that may better show the value of the genetic resources residing in botanic gardens.
Acknowledgments
Thank you to all the gardens (Birr, Lismore and Mount Usher) who gave samples for the analysis. Thanks also to Seamus O’Brien, Kilmacurragh for help in sourcing plant material. To Dr Peter Wyse Jackson for discussions on the project. Thank you to David Clarke, Archivist for additional information on the source of plant material in the National Botanic Gardens.
References
Charlesworth D. et Charlesworth B. Inbreeding depression and its evolutionary consequences // Annual Review of Ecology and Systematics. 1987. 18. 237—268.
Cicuzza D., Newton A. et Oldfield S. The Red List of Magnoliaceae // Flora and Fauna International (FFI) and Botanic Gardens International (BGCI). 2007. 52 p.
Forrest M. Magnolias in Ireland’s Gardens // Moorea 1984. 3. 31—34. 3 March 1984.
Forrest M. Trees and Shrubs Cultivated in Ireland. Kilkenny: Boethius Press, 1985.
IUCN. The IUCN Red List of Threatened Species, consulted 06.10.2015. http://www.iucnredlist.org/details/39011/0.
Jones C. J., Edwards K. J., Castaglione S. et al. Reproducibility testing of RAPD, AFLP and SSR markers in plants by a network of European laboratories // Molecular Breeding. 1997. 3. P. 381—390.
Kilpatrick J. Fathers of Botany: The Discovery of Chinese Plants by European Missionaries. Royal Botanic Gardens. 2014.
Kress W. J., Wurdack K. J., Zimmer E. A., Weigt L. A., Janzen D. H. Use of DNA barcodes to identify flowering plants // Proceedings of the National Academy of Sciences USA. 2005. 102. P. 8369—8374.
Maunder M., Higgens S. et Culham A. The effectiveness of botanic garden collections in supporting plant conservation: a European case study // Biodiversity et Conservation. 2001. 10. 3. P. 383—401.
Rivers M. C. et Wheeler L. Magnolia delavayi // The IUCN Red List of Threatened Species. 2014. e.T39011A2885519. http://dx.doi.org/10.2305.
Shaw J., Lickey E. B., Schilling E. E. et Small R. L. Comparison of whole chloroplast genome sequences to choose noncoding regions for phylogenetic studies in angiosperms: the tortoise and the hare III // American Journal of Botany. 2007. 94. P. 275—288.
Taberlet P., Gielly L., Pautou G. and Bouvet J. Universal primers for amplification of three noncoding regions of chloroplast DNA // Plant Molecular Biology. 1991. 17 P. 1105—1109.
Terachi T. Structural alterations of chloroplast genome and their significance to the higher plant evolution // Bulletin of the Institute for National Land Utilization Development. Kyoto Sangyo University. 1993. 14. P. 138—148. In Japanese.
Treseder N. G. Magnolias. London: Faber et Faber, 1978. 244 p.
Xun G., Quanan W., Yuanxue L. et Yanping Z. Pollination Biology of Cultivated Magnolia delavayi // Acta Botanica Yunnanica. 1998. 20. 1. P. 89—93. CBA: 535766.





